|
 |
| Korean J Anesthesiol > Volume 72(5); 2019 > Article |
|
Abstract
NOTES
1) Sample data (Survival2_PONV.csv) and the R console output of entire code are provided as supplemental information. Refer to online help or R statistical textbooks for detailed explanations of the argument. The included R code covers the process beginning with the survival analysis introduced in [1]. A detailed description of a violation of a proportional hazard assumption is provided in [14].
2) Terry M. survival: Survival Analysis. R package version 2.42-4. 2018. https://github.com/therneau/survival.
3) Alboukadel K, Marcin K, Przemyslaw B, Scheipl F. survminer: Drawing Survival Curves using 'ggplot2', R package version 0.4.3. 2018. http://www.sthda.com/english/rpkgs/survminer/.
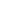
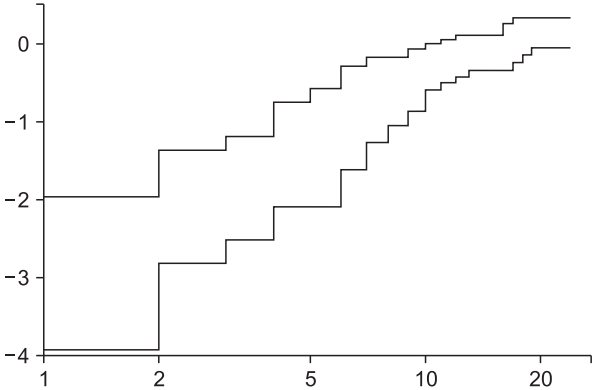
4) There are several ways to draw an LML plot in R; ‘plot.survfit’ with the argument ‘fun = “cloglog”’ provides an LML plot of the log-scaled x-axis. Most statistical references describe a log-scaled x-axis LML plot, whereas others describe a standard linear-scaled x-axis LML plot. The R code for a non-log-scaled LML plot can be created through the following. # Non-log scaled LML plot ponvsurv=Surv(PONV.raw$Time, PONV.raw$PONV) NLML.fun=function(p){return(log(-log(p)))} plot (survfit(ponvsurv ~ PONV.raw$Antiemetics), fun=NLML.fun)
5) R package ‘survival’ version 2.44-1 (updated in March 2019) has an error with an x value of 1 when log scaled using the x-axis. Versions before 2.44-1 work properly.
6) Schoenfeld residual is only for the patient who experienced the event. It is the difference between observed value of explanatory variable at a specific time and expected value of the explanatory variable (covariate) at a specific time which is a weighted-average value by likelihood of event from the risk set at that time point.
7) Some statistical software provides a method using scaled Schoenfeld residuals. Under a specific circumstance, these two results are different, although they mostly produce similar results. Please refer to the following: Grambsch, P.M. and Therneau, T.M. 1994. Proportional hazards tests and diagnostics based on weighted residuals. Biometrika 81: 515-526.
8) The command ‘ggadjustedcurves’ included in the ‘survminer’ library easily produces the survival curves of the CPH model. Unfortunately, this command still has minor functional errors such as in printing the 95% CI or labelling, and a somewhat complex ‘ggsurvplot’ is used in this example.
9) In R, categorical variables should be treated as a stratum for comparison using an LML plot of the CPH model.
10) Because various application methods and their variations are available, they are not discussed in detail herein.
11) This is a non-interaction stratified CPH model. Several survival functions are estimated through stratification, and if the explanatory variables have interactions with each other, the coefficients at each stratum may be different. In this case, it is assumed that an interaction model between explanatory variables and a likelihood ratio test provide clues to judge whether there is an interaction between explanatory variables. That is, if two or more variables are included in the model, it is necessary to check whether an interaction between them exists.
12) A clustered event time analysis and an accelerated failure time analysis are often applied to survival analysis methods in clinical study. A clustered event time analysis is similar with a stratified CPH model, and has certain advantages when each stratum has insufficient event cases. It has two types of processes, one is a marginal approach that estimates the survival function through an overall cluster from the pooled effect of each stratum, and another is a conditional approach that estimates the survival function from the heterogeneity between clusters. An accelerated failure time analysis estimates the model similarly with a linear regression based on a Weibull distribution or log-logistic distribution. Unlike a CPH model that continuously maintains the risk ratio of the covariates, this model assumes that the disease process can be accelerated or decelerated over time.
Supplementary Materials
Fig. 1.
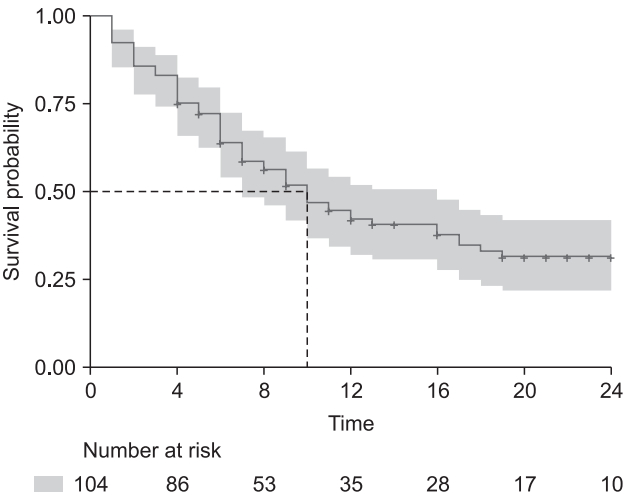
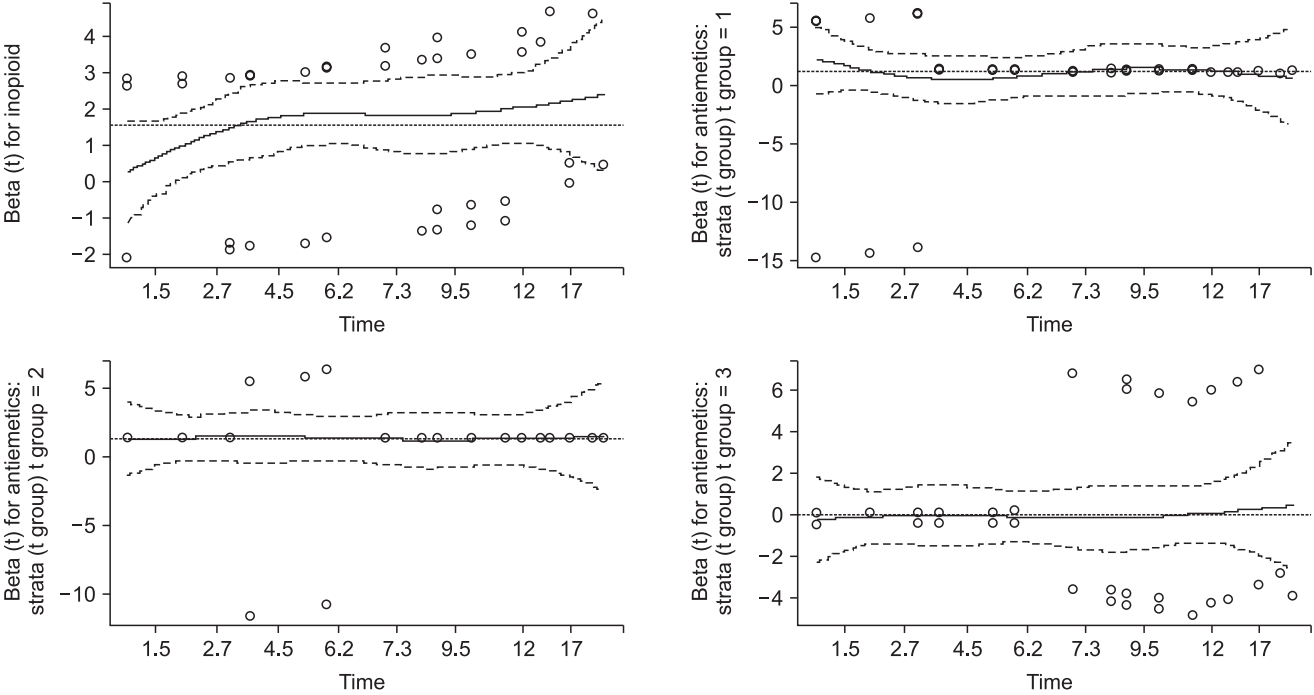
Fig. 2.
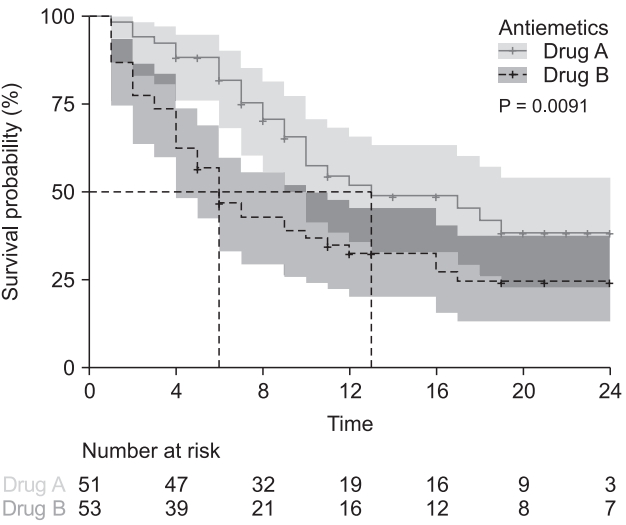
Fig. 3.

Fig. 4.

Fig. 6.
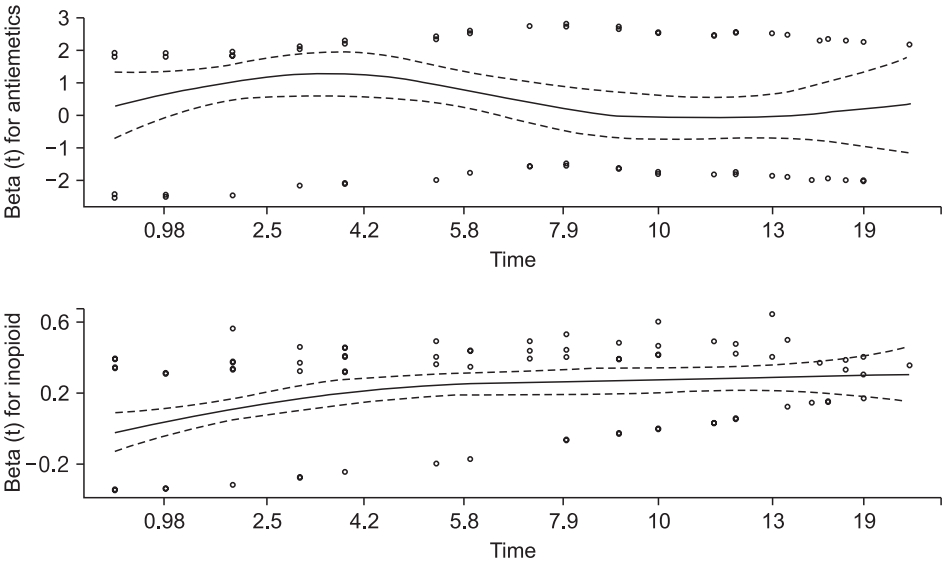
Fig. 7.
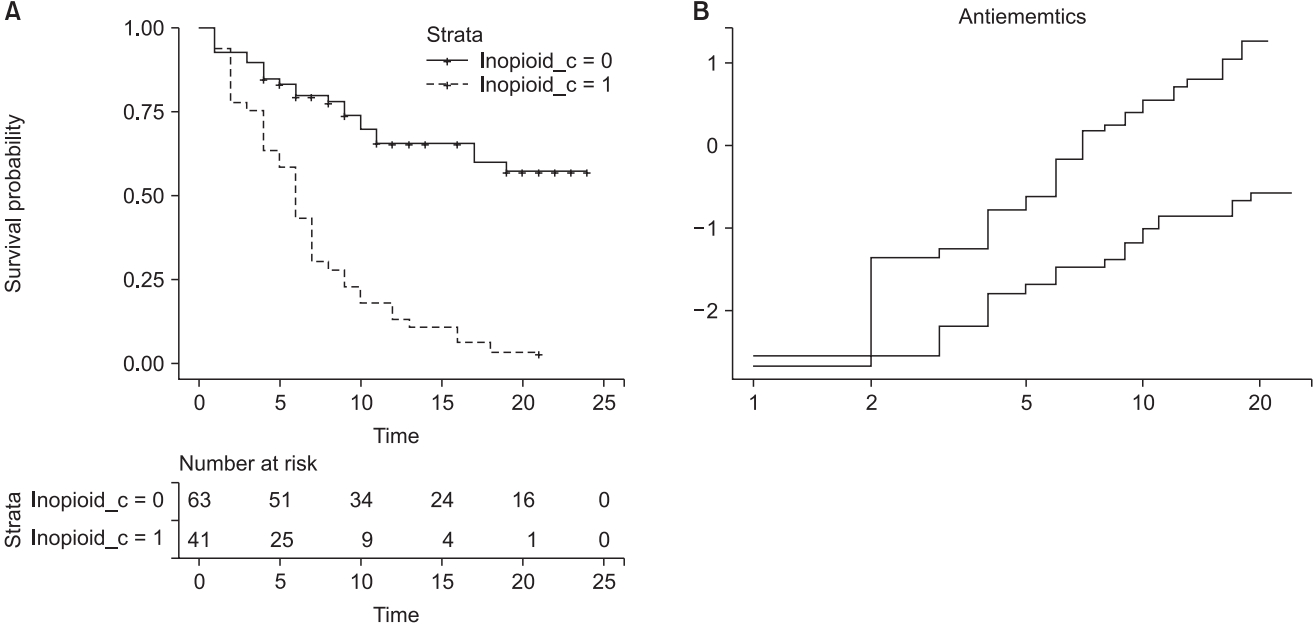
Fig. 8.
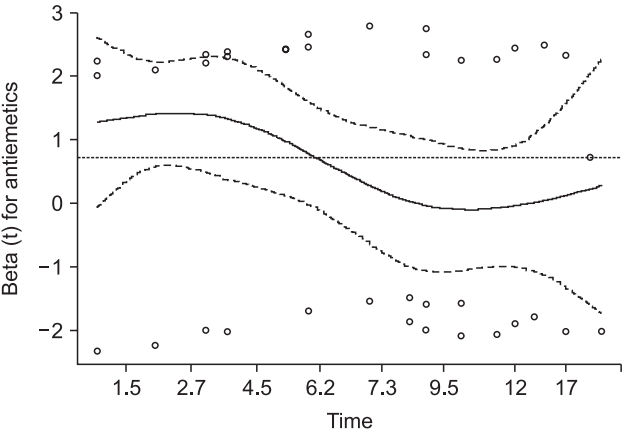
Fig. 10.

Fig. 11.

Table 1.
|
Result of command “head(PONV.raw) |
|||||||
|---|---|---|---|---|---|---|---|
| No | Antiemetics | Age | Wt | Inopioid | Time | PONV | |
| 1 | 1 | 0 | 48 | 78.5 | 0 | 4 | 0 |
| 2 | 3 | 0 | 54 | 88.3 | 100 | 21 | 0 |
| 3 | 4 | 0 | 22 | 49.4 | 0 | 14 | 0 |
| ︙ | ︙ | ︙ | ︙ | ︙ | ︙ | ︙ | ︙ |
From the left, each column contains each coded variable: The first column has a number automatically generated by R, variable ‘No’ is a coded number in the original data, ‘Antiemetics’ has a value of 0 for Drug A and 1 for Drug B, ‘Age’ and ‘Wt’ are the actual patients’ age and body weight, ‘Inopioid’ is the amount of opioid used during surgery, ‘Time’ indicates the onset time of postoperative nausea and vomiting (PONV), and ‘PONV’ is coded as 1 when the patient experienced PONV.
Table 2.
n: total number of cases, Events: number of patients who experienced PONV, Median: median survival time, 0.95LCL: lower limit of 95% confidence interval, 0.95UCL: upper limit of 95% confidence interval, n.risk: number at risk, n.event: number of event, Survival: survival rate, std.err: standard error of survival rate, Lower/upper 95% CI: lower/upper limits of 95% confidence interval.
Table 3.
Antiemetics = 0 and 1 indicate Drugs A and B, respectively. Because the variable ‘Antiemetics’ is coded as 0 for drug A and 1 for drug B, the R output only describes these as ‘Antiemetics = 0 and 1’. n: total number of cases, Events: number of patients who experienced postoperative nausea and vomiting, Median: Median survival time, 0.95LCL: lower limit of 95% confidence interval, 0.95UCL: upper limit of 95% confidence interval, n.risk: number at risk, n.event: number of events, Survival: survival rate, std.err: standard error of survival rate, Lower/upper 95% CI: lower/upper limits of 95% confidence interval, Chisq: chi-squared statistics.
Table 4.
Table 5.
‘Antiemetics’ is coded as 0 for Drug A or 1 for Drug B in the original data. ‘Inopioid’ is the amount of opioid used during surgery. coef: the value of coefficient, exp(coef): exponential value of coefficient, se(coef): standard error of coefficient, z: z-statistics, Pr(>|z|): P value of given z-statistics, Signif. codes: codes for significance marking.
Table 6.
|
Results of ‘print(sf.residual)’ |
|||
|---|---|---|---|
| rho | chisq | P | |
| Antiemetics | −0.275 | 4.5 | 0.0340 |
| Inopioid | 0.307 | 5.36 | 0.0206 |
| GLOBAL | NA | 10.26 | 0.0059 |
Table 7.
|
Results of ‘head (PONV.raw)’ |
|||||||||
|---|---|---|---|---|---|---|---|---|---|
| No | Antiemetics | Age | Wt | Inopioid | Time | PONV | Survobj | Inopioid_c | |
| 1 | 1 | 0 | 48 | 78.5 | 0 | 4 | 0 | 4+ | 0 |
| 2 | 3 | 0 | 54 | 88.3 | 100 | 21 | 0 | 21+ | 1 |
| 3 | 4 | 0 | 22 | 49.4 | 0 | 14 | 0 | 14+ | 0 |
| ︙ | ︙ | ︙ | ︙ | ︙ | ︙ | ︙ | ︙ | ︙ | ︙ |
‘Survobj’ is a variable created by an R command during the process of a Kaplan–Meier estimate, and indicates a survival object. ‘Inopioid_c’ is a newly created categorical variable based on ‘Inopioid’, which is coded as 0 for an opioid not used or 1 for an opioid used during operation. From left, each column contains each coded variable: The first column has a number automatically generated by R, the variable ‘No’ is a coded number in the original data, ‘Antiemetics’ has a value of 0 for Drug A and 1 for Drug B, ‘Age’ and ‘Wt’ are the actual patients’ age and body weight, ‘Inopioid’ is the amount of opioid used during surgery, ‘Time’ indicates the onset time of postoperative nausea and vomiting, and ‘PONV’ is coded as 1 when the patient experienced postoperative nausea and vomiting.
Table 8.
‘Antiemetics’ is coded as 0 for Drug A or 1 for Drug B in the original data. ‘Inopioid_c’ is a newly created categorical variable based on ‘Inopioid’, which is coded as 0 for an opioid not used or 1 for an opioid used during operation. coef: the value of coefficient, exp(coef): exponential value of coefficient, se(coef): standard error of coefficient, z: z-statistics, Pr(>|z|): P value of given z-statistics, Signif. codes: codes for significance marking.
Table 9.
All personal data are separated according to a preset time period (at 3 and 6 h). The same ‘id’ number indicates the same person. For example, data with id = 1 are separated into two time periods. The first period starts from time = 0 (tstart = 0) and ends at 3 (Time = 3) and PONV does not occur. The second period starts from 3 to 4 (the observation is prematurely ended before 6) and PONV does not occur. The same time period is indicated as tgroup (time group) in the last column. The other variables are the same as in Table 7.
Table 10.
‘Antiemetics’ is coded as 0 for Drug A or 1 for Drug B in the original data. ‘Inopioid’ is the amount of opioid used during surgery. The split time periods are presented as Antiemetics:strata(tgroup)tgroup = 1 for the time period from 0 to 3, Antiemetics:strata(tgroup)tgroup = 2 for the time period from 3 to 6, and Antiemetics:strata(tgroup)tgroup = 3 for the time period from 6 to the end of the observation. coef: the value of coefficient, exp(coef): exponential value of coefficient, se(coef): standard error of coefficient, z: z-statistics, Pr(>|z|): P value of given z-statistics, Signif. codes: codes for significance marking.
Table 11.
Proportional hazard assumed Kaplan–Meier analysis results are presented in the upper part of the table. Note that this result is the same as in Table 3. The lower part of this table presents the results of a Cox regression with a time-dependent coefficient. The median survival is different from the proportional hazard assumed analysis. Antiemetics = 0 and 1 indicate Drugs A and B respectively. n: total number of cases, Events: number of patients who experienced postoperative nausea and vomiting, 0.95LCL: lower limit of 95% confidence interval, or 0.95UCL: upper limit of 95% confidence interval.